Biology, Free Full-Text
Por um escritor misterioso
Descrição
Microbial symbionts range from mutualistic to commensal to antagonistic. While these roles are distinct in their outcome, they are also fluid in a changing environment. Here, we used the Lotus japonicus–Mesorhizobium japonicum symbiosis to investigate short-term and long-term shifts in population abundance using an effective, fast, and low-cost tracking methodology for M. japonicum. We use quantitative polymerase chain reaction (qPCR) to track previously generated signature-tagged M. japonicum mutants targeting the Tn5 transposon insertion and the flanking gene. We used a highly beneficial wild type and moderately beneficial and non-beneficial mutants of M. japonicum sp. nov. to demonstrate the specificity of these primers to estimate the relative abundance of each genotype within individual nodules and after serial transfers to new hosts. For the moderate and non-beneficial genotypes, qPCR allowed us to differentiate genotypes that are phenotypically indistinguishable and investigate host control with suboptimal symbionts. We consistently found the wild type increasing in the proportion of the population, but our data suggest a potential reproductive trade-off between the moderate and non-beneficial genotypes. The multi-generation framework we used, coupled with qPCR, can easily be scaled up to track dozens of M. japonicum mutants simultaneously. Moreover, these mutants can be used to explore M. japonicum genotype abundance in the presence of a complex soil community.

Stream [EBOOK] 📕 Princeton Review AP Biology Premium Prep, 2023
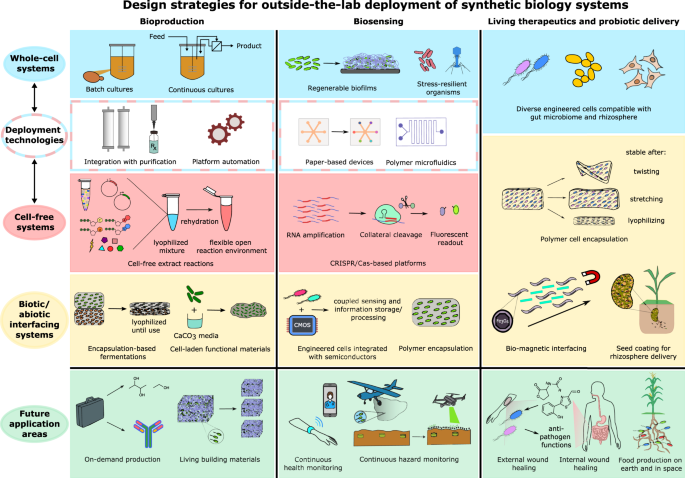
Applications, challenges, and needs for employing synthetic
This book is for your convenience. It is available for printing from our site. This book contains the sheets you would print if using the online

EP Biology Printables: Levels 1-4: Part of the Easy Peasy All-in-One Homeschool

AP Biology (Paperback)
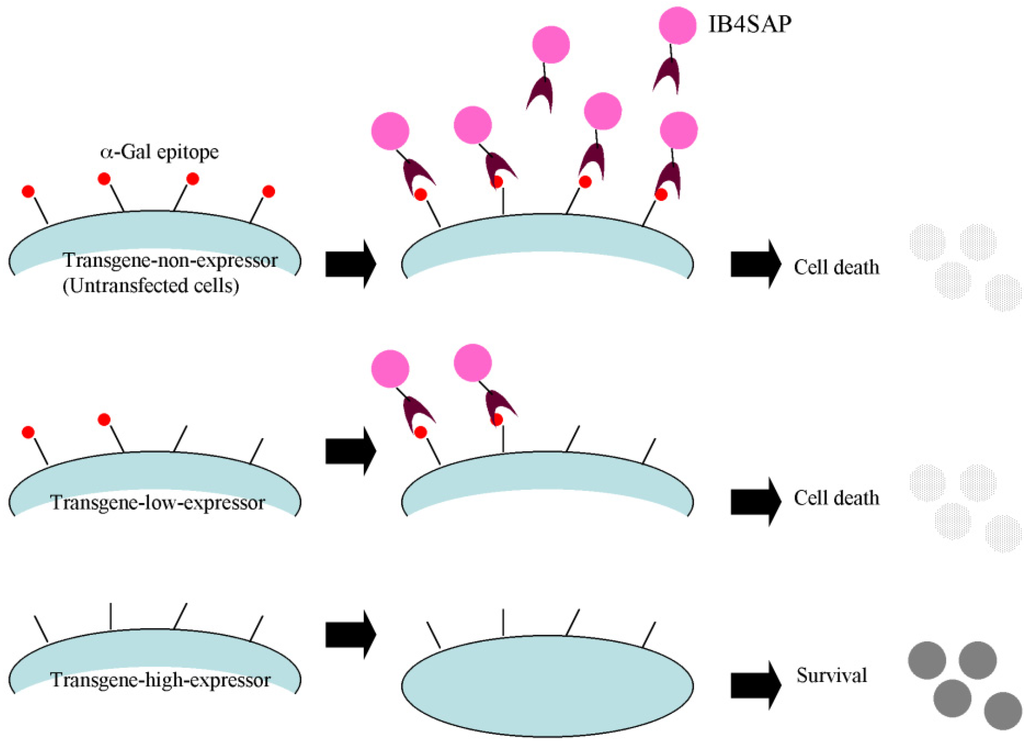
Biology, Free Full-Text

Full text access through Strategian - Strategian Science
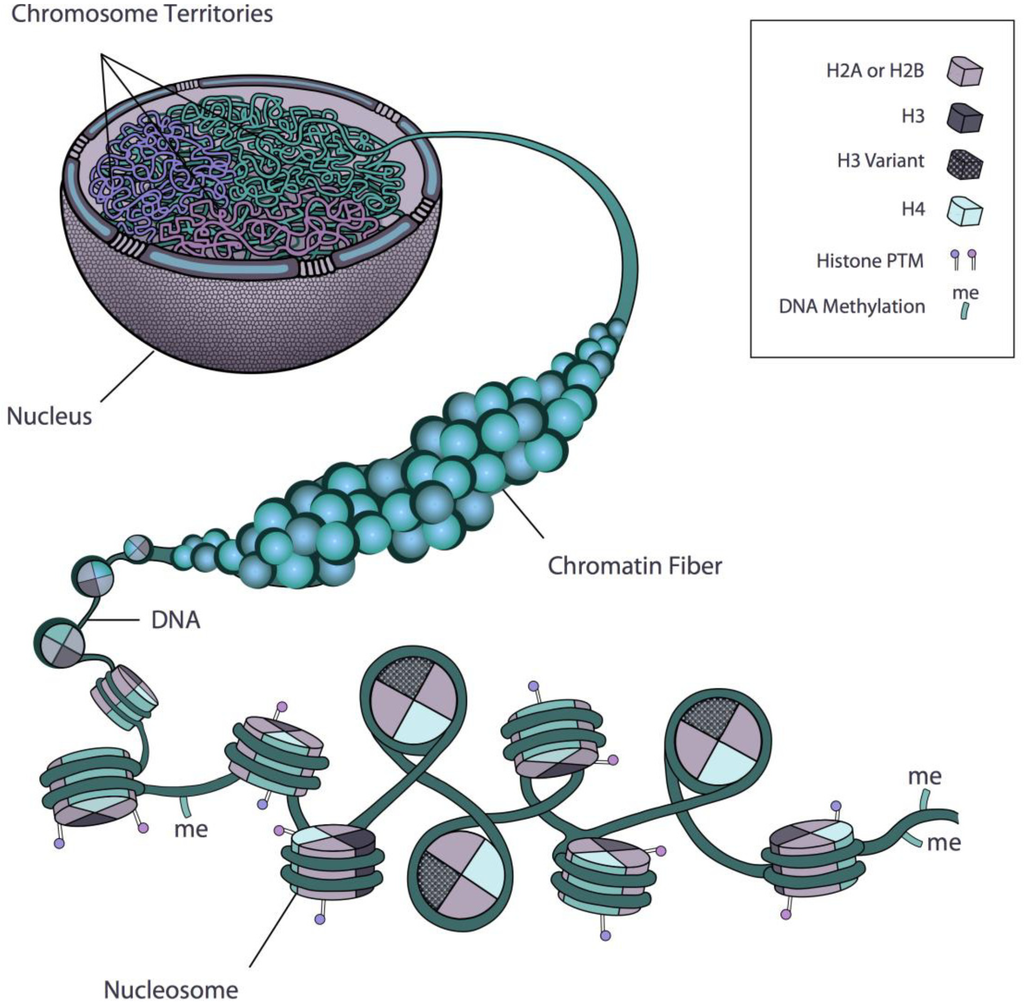
Biology, Free Full-Text

Journal of Biological Rhythms: Sage Journals

Cell-free biology: exploiting the interface between synthetic
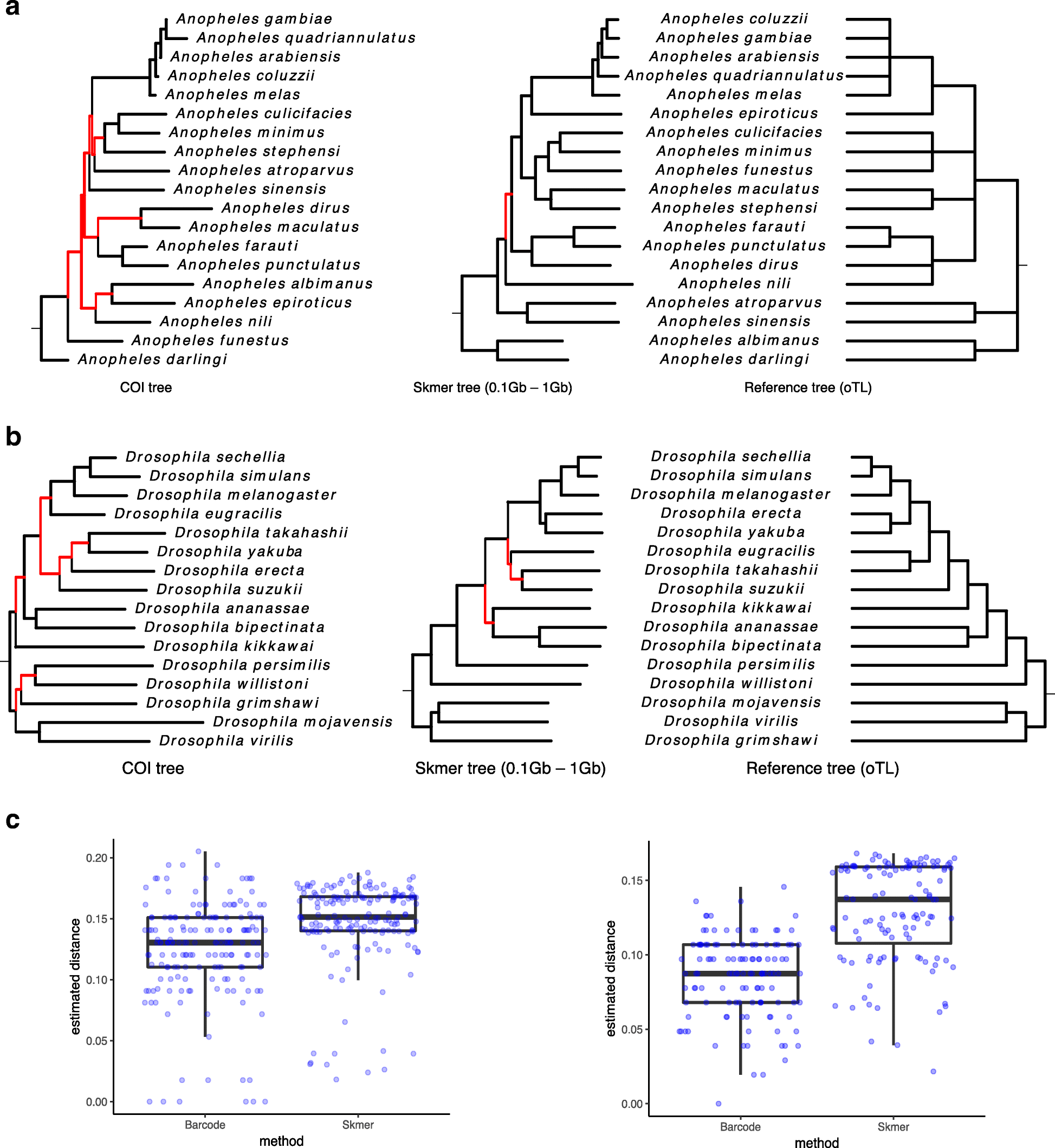
Skmer: assembly-free and alignment-free sample identification







